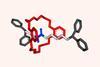
A team surrounding Ivan Huc, from Ludwig-Maximilians-University of Munich in Germany, has made a 12.9kDa pseudo-quaternary structure composed of four aromatic oligoamide helix-turn-helix modules.1 Single crystal x-ray diffraction analysis of the abiotic foldamer shows some previously identified assembly patterns, as well as new ones involving multiple water molecule bridges. ‘This contribution to foldamer research pushes the complexity limit of artificial assemblies,’ remarks Helma Wennemers, a chemical biology expert from ETH Zurich in Switzerland, who was not involved in the project.
The structure of natural proteins can be described using four levels of complexity – from primary to quaternary. It’s particularly important for proteins to fold into precise tertiary and quaternary structures to function correctly. Abiotic foldamers, like their biological counterparts, adopt a conformationally ordered state in solution, yet are based on building blocks not seen in nature. In these systems, sophisticated functions are also predicted to arise at higher levels of structural complexity. Samuel Gellman, an expert in foldamers at the University of Wisconsin–Madison, US, explains that the design, synthesis and characterisation of unnatural oligomers with higher-order structures reminiscent of those found among proteins and nucleic acids is an ambitious and widely pursued goal in the chemistry community. ‘Ultimately, chemists hope to endow these foldamer systems with powerful functions analogous to those of natural enzymes or receptors, or even functions that are not possible for biopolymers.’
Foldamer research has already shown that different abiotic backbones can fold into helices. Aromatic oligoamides that possess hydrogen-bonding functionalities and units capable of aromatic stacking are one such class that can form helical conformations in organic solvents. In 2018, another team surrounding Huc designed and made the first true abiotic tertiary foldamer structure, which was based on an aromatic oligoamide helix-turn-helix motif.2
The next milestone
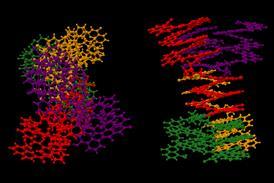
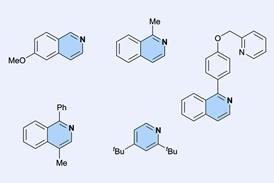
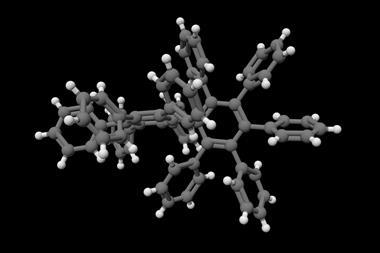
Building on this work, Huc and colleagues have now experimented with various helix-turn-helix structures incorporating a di-, tri-, or tetra-ethylene glycol flexible linker, as well as hydrogen bond donor and acceptor groups. The handedness of the helices was controlled through the presence, or absence, of chiral components based on 2-(2-aminophenoxy)-propionic acid or a (1H)-camphanyl group. Huc’s team discovered a case where four helix-turn-helix modules based on the same sequence and lacking any chiral component self-assembled to form an architecture composed of eight helices. The helices were arranged in three distinct domains, with several of the helix-to-helix hydrogen bonds mediated by bridging water molecules. It had a quaternary-like structure that ‘was impossible to predict,’ comments Huc. He explains that the work ‘is about transitioning from synthetic models that just mimic a secondary fold, like a helix or a linear strand, to structures that are really closer to what proteins are like, with more complex tertiary folds that are composed of multiple helices, or multiple secondary forms.’
Sarah Pike from the University of Birmingham, UK, who researches synthetic helical foldamers, notes that ‘creating such bioinspired architectures is important as these synthetic scaffolds have widespread applications both in biological and artificial settings. The identification of an abiotic eight-helix bundle serves to highlight the immense potential of designed foldamers to probe new chemical space.’
With a view to the future, Huc says that ‘there’s still a stage that we haven’t reached, which would be getting tertiary structures that really have the sophistication of a small protein. We need to elaborate the sophistication a bit more and then we can think of trying to implement functions.’ Automatic synthesisers will also play a role, adds Huc, as ‘now that a machine can do it, it saves so much time and is very efficient. We’re not frightened by the size of the objects we can synthesise and it’s becoming relatively easy to be in the 10kDa range by synthesis, so like small proteins.’
‘This study represents a huge step forward towards enabling the controlled construction of biomimetic self-assembled architectures from abiotic foldamers, with important and far-reaching implications on the future design of functional foldamers,’ says Pike. ‘[It] will enable access to novel, abiotic foldamer scaffolds that possess complex topologies to rival that of biological systems.’
References
1 F S Menke et al, Chem. Sci., 2023, DOI: 10.1039/d3sc00267e
2 S De et al, Nat. Chem., 2018, 10, 51 (DOI: 10.1038/nchem.2854)














No comments yet