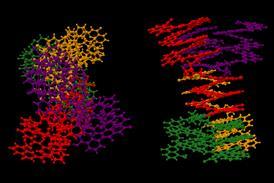
Exploring AlphaFold’s knowledge of energy landscapes

The algorithm needs a little help to find the global energy minimum
The success of AlphaFold for predicting the structures of more than 200 million proteins, announced last August, led to excited claims that the algorithm will revolutionise biology, drug discovery and molecular medicine. That remains to be seen, but some were keen to temper the hype by pointing out that AlphaFold had not in fact ‘solved the protein-folding problem’. Rather, it had sidestepped the question by using machine learning to find associations between sequence and known structures that it then generalised to unknown structures.
Unlike, say, a molecular dynamics simulation, AlphaFold – devised by a team at DeepMind, an offshoot of Google – doesn’t attempt to recapitulate the molecular pathway leading to the folded structure. It just uses the correlations it has learnt between sequence and chain shape. To identify these for an arbitrary primary amino-acid sequence, however, the algorithm needs to compile collections of sequences that are closely related to the target sequence, called multiple sequence alignments (MSAs). These give the algorithm a sense of what the consequences are of amino-acid substitutions in this part of configuration space. This requirement for MSAs is a problem if there are few known proteins that are close homologues to the target.